ABOUT
LinkedOmics is publicly available portal that includes multi-omics data from all 32 TCGA Cancer types and 10 Clinical Proteomics Tumor Analysis Consortium (CPTAC) cancer cohorts.
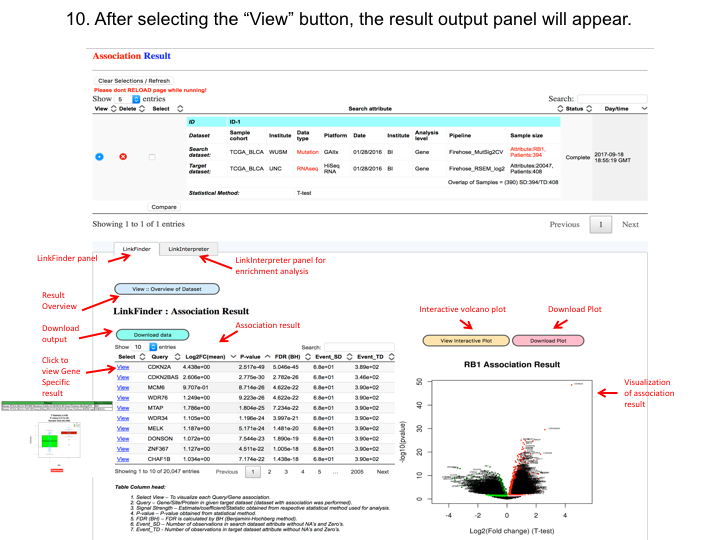
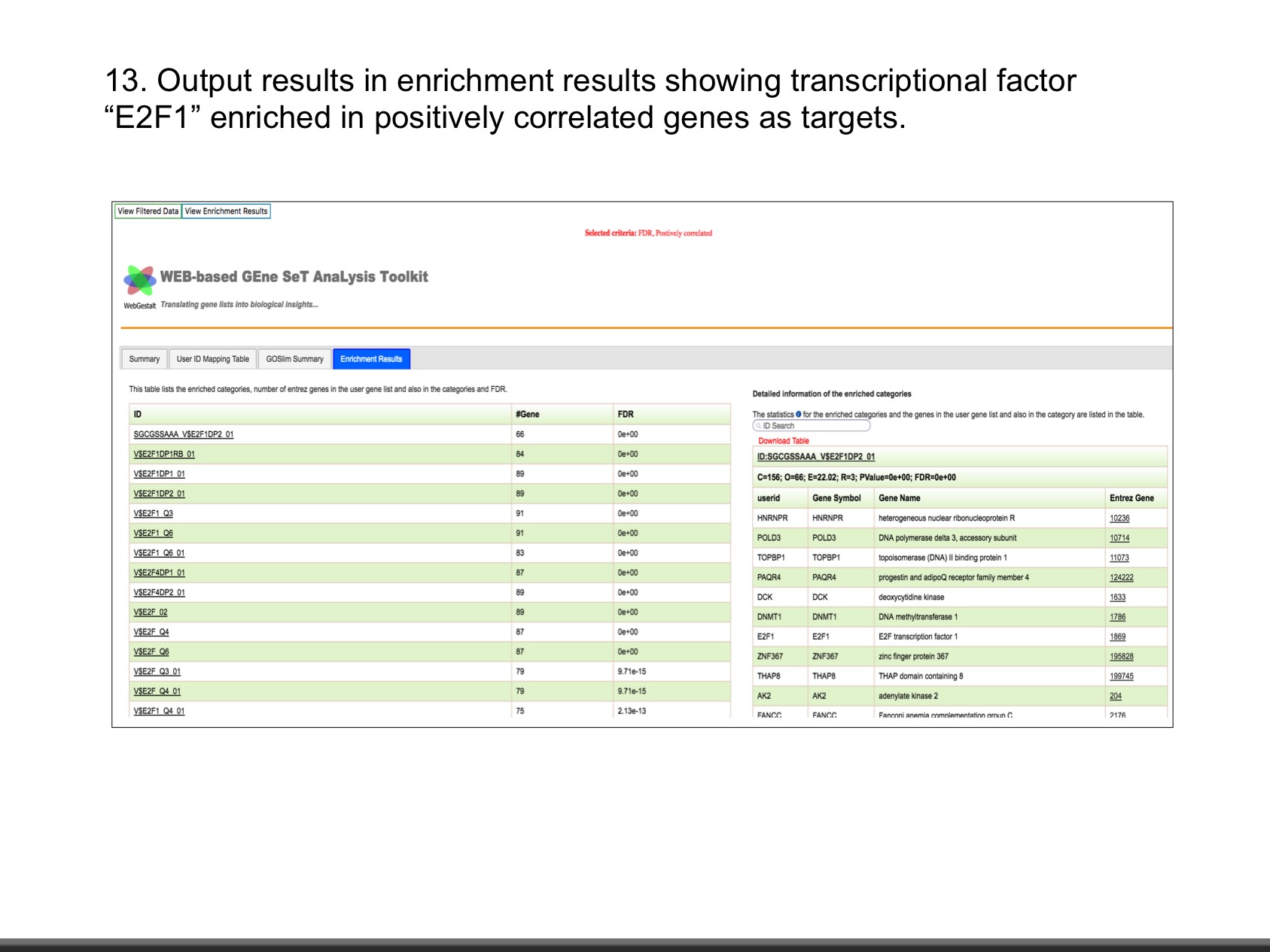
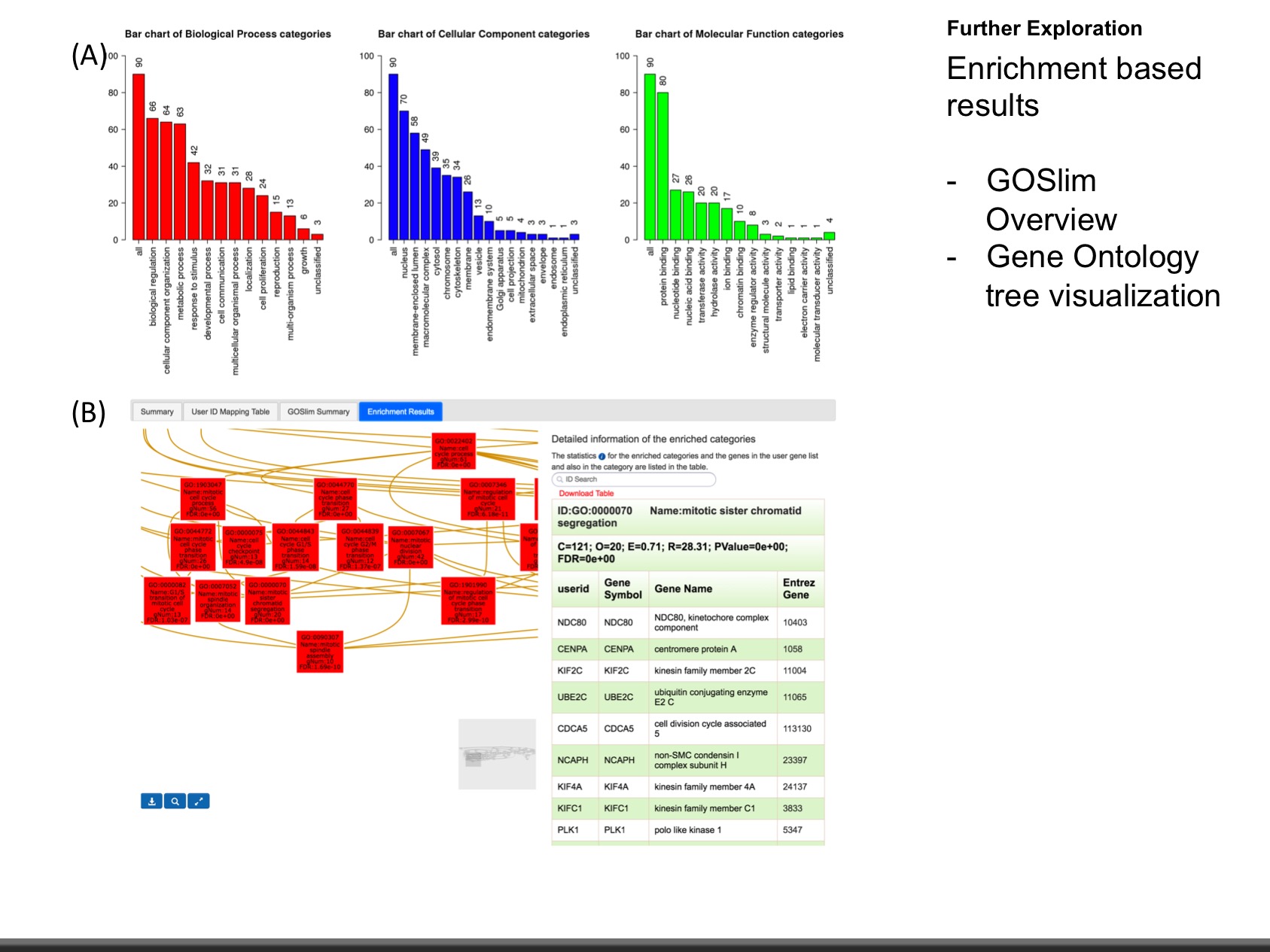
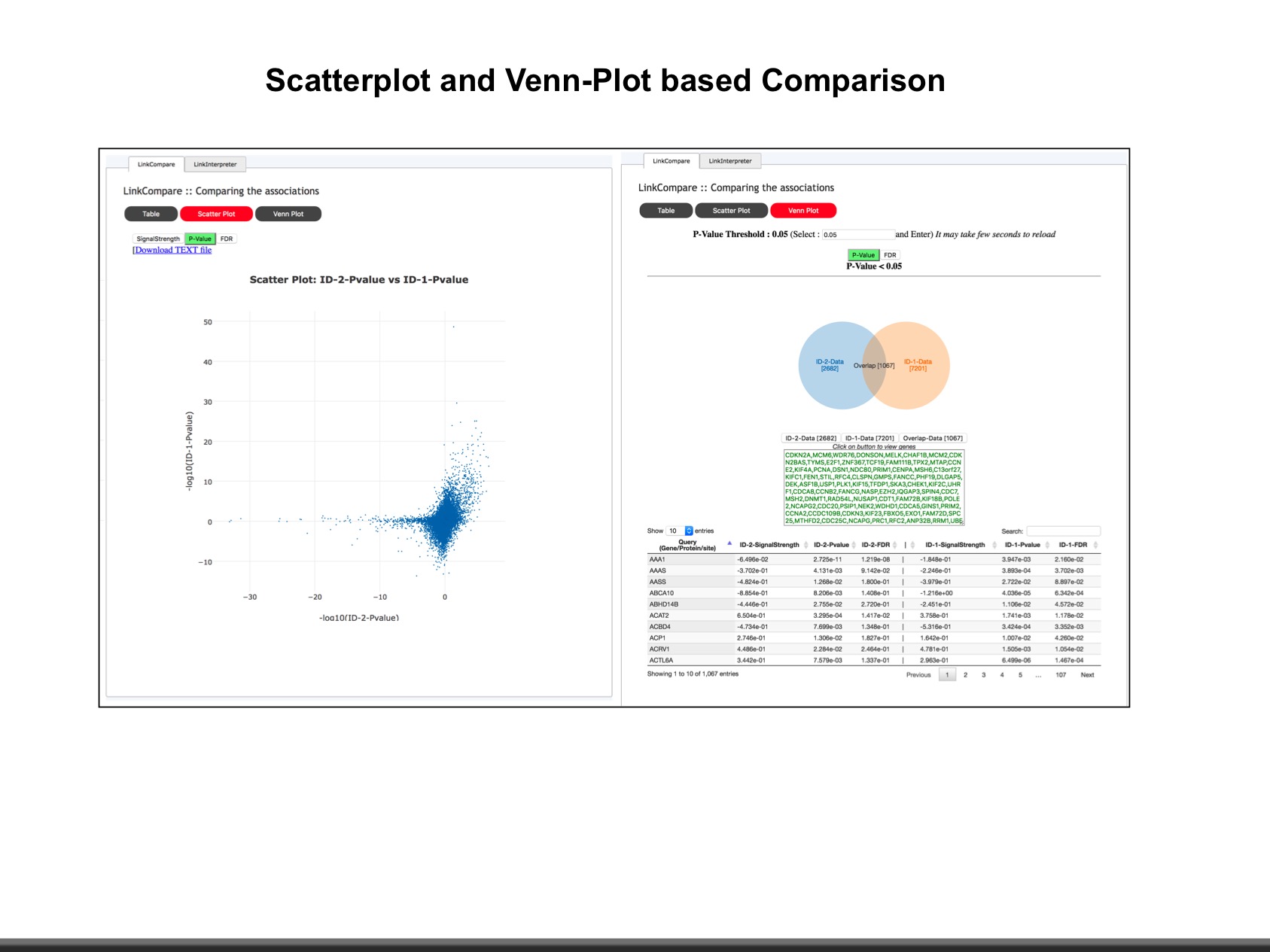
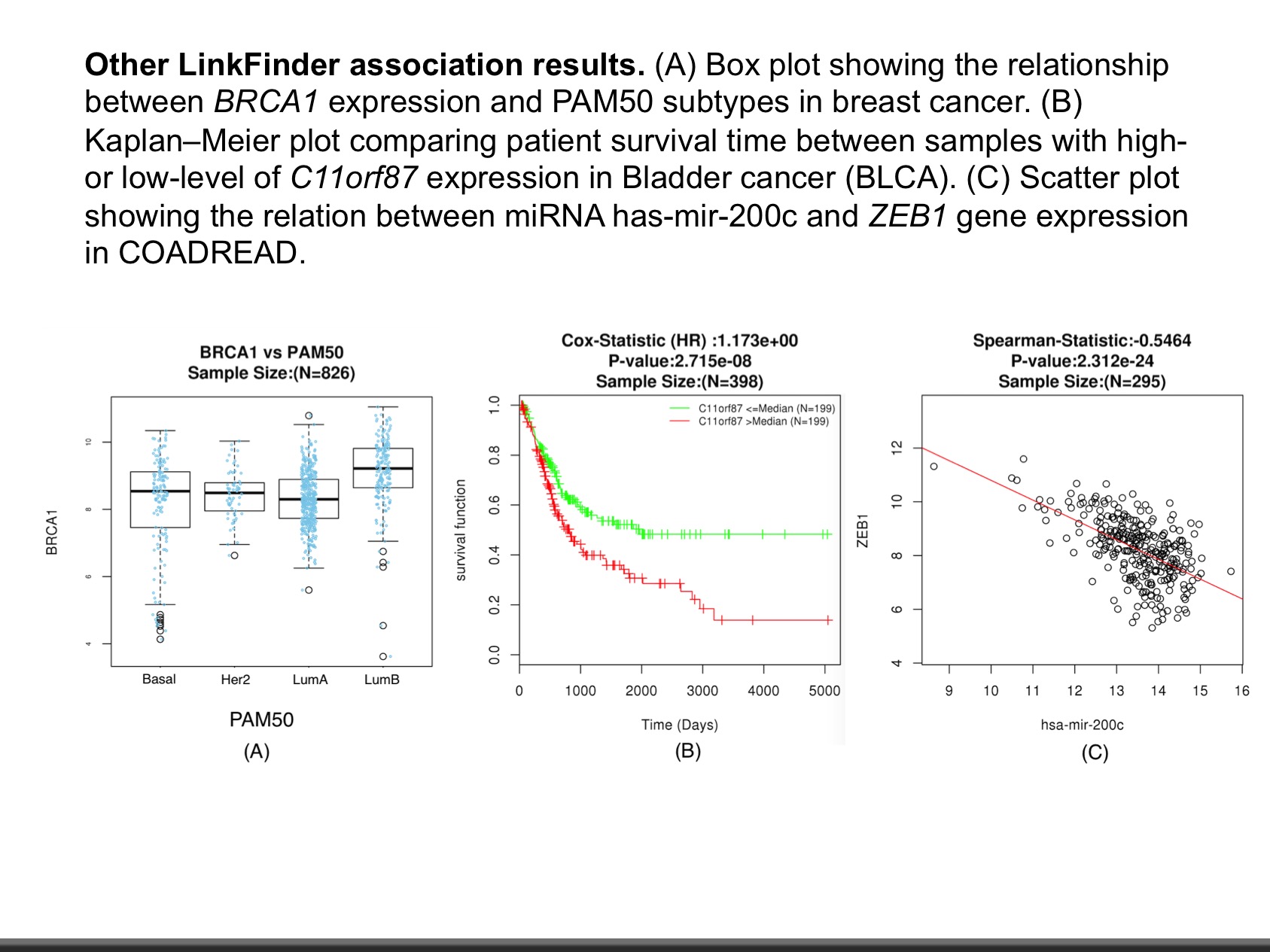
The web application has three analytical modules: LinkFinder, LinkInterpreter and LinkCompare. LinkFinder allows users to search for attributes that are associated with a query attribute, such as mRNA or protein expression signatures of genomic alterations, candidate biomarkers of clinical attributes, and candidate target genes of transcriptional factors, microRNAs, or protein kinases. Analysis results can be visualized by scatter plots, box plots, or Kaplan-Meier plots. To derive biological insights from the association results, the LinkInterpreter module performs enrichment analysis based on Gene Ontology, biological pathways, network modules, among other functional categories. The LinkCompare module uses visualization functions (interactive venn diagram, scatter plot, and sortable heat map) and meta-analysis to compare and integrate association results generated by the LinkFinder module, which supports multi-omics analysis in a cancer type or pan-cancer analysis.
LinkedOmics provides a unique platform for biologists and clinicians to access, analyze and compare cancer multi-omics data within and across tumor types.
The web application has three analytical modules: LinkFinder, LinkInterpreter and LinkCompare. LinkFinder allows users to search for attributes that are associated with a query attribute, such as mRNA or protein expression signatures of genomic alterations, candidate biomarkers of clinical attributes, and candidate target genes of transcriptional factors, microRNAs, or protein kinases. Analysis results can be visualized by scatter plots, box plots, or Kaplan-Meier plots. To derive biological insights from the association results, the LinkInterpreter module performs enrichment analysis based on Gene Ontology, biological pathways, network modules, among other functional categories. The LinkCompare module uses visualization functions (interactive venn diagram, scatter plot, and sortable heat map) and meta-analysis to compare and integrate association results generated by the LinkFinder module, which supports multi-omics analysis in a cancer type or pan-cancer analysis.
LinkedOmics provides a unique platform for biologists and clinicians to access, analyze and compare cancer multi-omics data within and across tumor types.
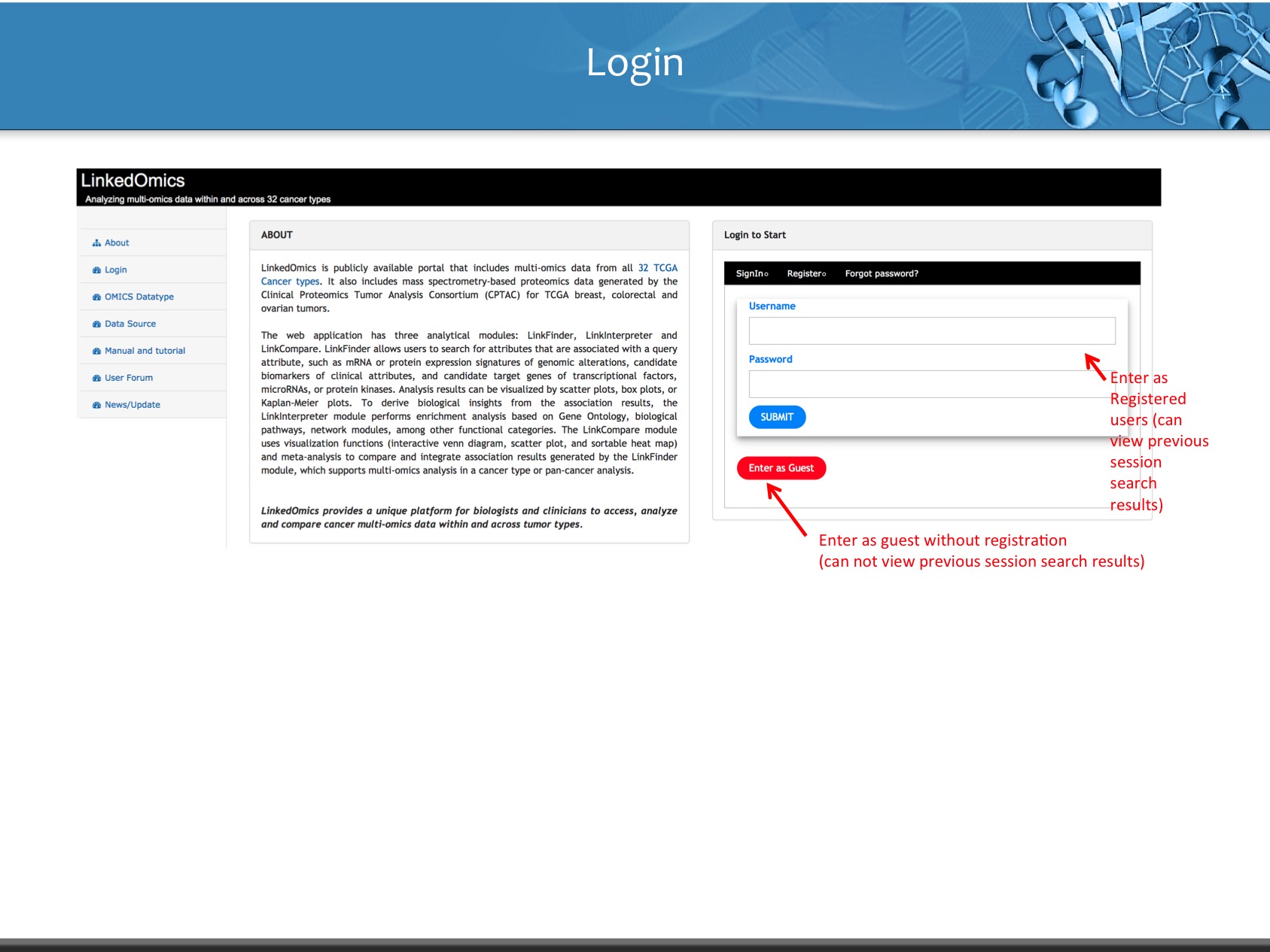
Login
Please cite this article as,
-
Suhas V Vasaikar, Peter Straub, Jing Wang, Bing Zhang, LinkedOmics: analyzing multi-omics data within and across 32 cancer types, Nucleic Acids Research, Volume 46, Issue D1, 4 January 2018, Pages D956–D963.https://doi.org/10.1093/nar/gkx1090
- Clinical Data : It inculdes attributes like age, overall survival, phological stage (I, II, III, IV), TNM staging, Clinical subtype, Molecular Subtype, number of lymph nodes, radiation therapy.
- Copy Number (Level: Focal, Gene) : Normalized copy number (SNPs) ans Copy number alterations for aggregated/segmented regions, per sample
- miRNA (Leve: Gene, Isoform) : Normalized signals per probe or probe set for each participant's tumor sample
- Mutation (Level: Site, Gene) : Mutation calls for each participant
- Methylation (Level: Gene) : Calculated beta values mapped to genome, per sample
- RNAseq (Level: Gene) : The normalized expression signal of individual Gene (transcripts), per sample
- RPPA (Level: Analyte, Gene) : Normalized protein expression for each gene, per sample
- Proteomics (Level: Gene) : Average log-ratio of sample reporter-ion to common reference of peptide ions associated with the gene in acquisitions from a specific biological Sample (Unshared Log Ratio-Average log-ratio of sample reporter-ion to common reference of peptide ions of unshared peptides only associated with the gene in acquisitions from a specific biological sample).
- Phospho-Proteomics (Level: Site) : Average log-ratio of sample reporter-ion to common reference of peptide ions associated with phosphorylated site combinations in acquisitions from a specific biological sample (CDAP Protein Report).
- Glyco-Proteomics (Level: Site) : Average log-ratio of sample reporter-ion to common reference of peptide ions associated with deglycosylated N-glycosylation site combinations in acquisitions from a specific biological sample (CDAP Protein Report).
For more information (Click here)
| Cancer Type | Cohort Source | Cancer ID | Samples | Permission | Link | Data Download |
|---|---|---|---|---|---|---|
| Pediatric Neuroblastoma | Pediatric | Neuroblastoma | 344 | Y | cBioPortal |
Download
|
| CPTAC-ICPC - Combined Lung Adenocarcinoma Cohorts | CPTAC-ICPC | LUAD | 794 | Y | CPTAC, Publication |
Download
|
| Paediatric High Grade Glioma | Pediatric | HGG | 86 | Y | cBioPortal, Publication |
Download
|
| Prostate adenocarcinoma | TCGA | PRAD | 499 | Y | TCGA, GDAC |
Download
|
| Sarcoma | TCGA | SARC | 261 | Y | TCGA, GDAC |
Download
|
| Skin Cutaneous Melanoma | TCGA | SKCM | 470 | Y | TCGA, GDAC |
Download
|
| Pheochromocytoma and Paraganglioma | TCGA | PCPG | 179 | Y | TCGA, GDAC |
Download
|
| Ovarian serous cystadenocarcinoma | TCGA | OV | 591 | Y | TCGA, GDAC, CPTAC |
Download
|
| Mesothelioma | TCGA | MESO | 87 | Y | TCGA, GDAC |
Download
|
| Stomach adenocarcinoma | TCGA | STAD | 443 | Y | TCGA, GDAC |
Download
|
| Pancreatic adenocarcinoma | TCGA | PAAD | 185 | Y | TCGA, GDAC |
Download
|
| Testicular Germ Cell Tumors | TCGA | TGCT | 134 | Y | TCGA, GDAC |
Download
|
| Uterine Carcinosarcoma | TCGA | UCS | 57 | Y | TCGA, GDAC |
Download
|
| Adrenocortical carcinoma | TCGA | ACC | 92 | Y | TCGA, GDAC |
Download
|
| Uterine Corpus Endometrial Carcinoma | TCGA | UCEC | 548 | Y | TCGA, GDAC |
Download
|
| Thymoma | TCGA | THYM | 124 | Y | TCGA, GDAC |
Download
|
| Lung squamous cell carcinoma | TCGA | LUSC | 504 | Y | TCGA, GDAC |
Download
|
| Thyroid carcinoma | TCGA | THCA | 503 | Y | TCGA, GDAC |
Download
|
| Stomach and Esophageal carcinoma | TCGA | STES | 628 | Y | TCGA, GDAC |
Download
|
| Uveal Melanoma | TCGA | UVM | 80 | Y | TCGA, GDAC |
Download
|
| Colorectal adenocarcinoma | TCGA | COADREAD | 629 | Y | GDAC, CPTAC |
Download
|
| Esophageal carcinoma | TCGA | ESCA | 185 | Y | TCGA, GDAC |
Download
|
| Glioblastoma multiforme | TCGA | GBM | 595 | Y | TCGA, GDAC |
Download
|
| Cholangiocarcinoma | TCGA | CHOL | 45 | Y | TCGA, GDAC |
Download
|
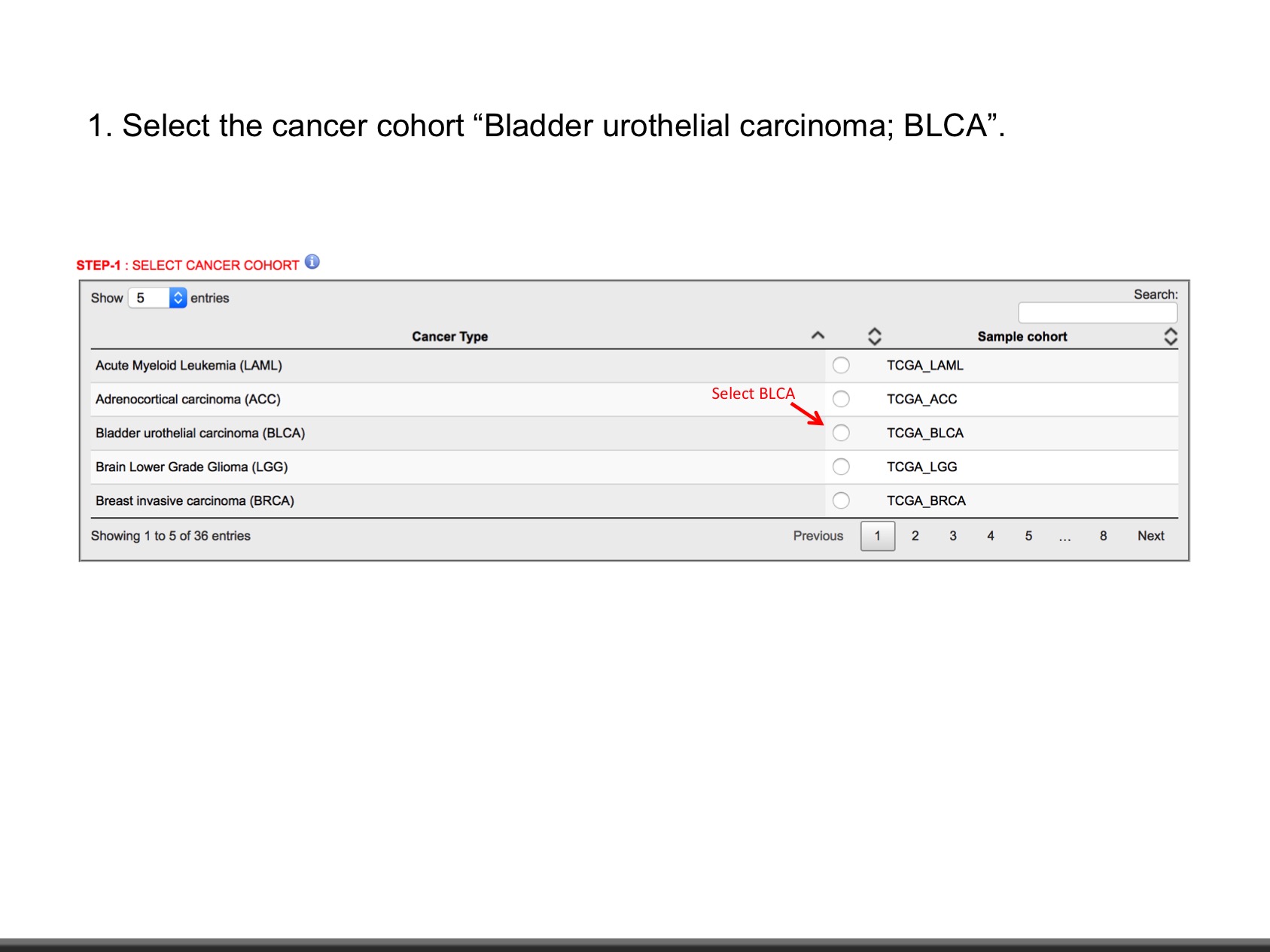
| Bladder urothelial carcinoma | TCGA | BLCA | 412 | Y | TCGA, GDAC |
Download
|
| Lung adenocarcinoma | TCGA | LUAD | 522 | Y | TCGA, GDAC |
Download
|
| Cervical and endocervical cancers | TCGA | CESC | 307 | Y | TCGA, GDAC |
Download
|
| Breast invasive carcinoma | TCGA | BRCA | 1097 | Y | TCGA, GDAC, CPTAC |
Download
|
| Glioma | TCGA | GBMLGG | 1110 | Y | TCGA, GDAC |
Download
|
| Lymphoid Neoplasm Diffuse Large B-cell Lymphoma | TCGA | DLBC | 48 | Y | TCGA, GDAC |
Download
|
| Acute Myeloid Leukemia | TCGA | LAML | 200 | Y | TCGA, GDAC |
Download
|
| Head and Neck squamous cell carcinoma | TCGA | HNSC | 528 | Y | TCGA, GDAC |
Download
|
| Liver hepatocellular carcinoma | TCGA | LIHC | 377 | Y | TCGA, GDAC |
Download
|
| Brain Lower Grade Glioma | TCGA | LGG | 515 | Y | TCGA, GDAC |
Download
|
| Kidney renal papillary cell carcinoma | TCGA | KIRP | 291 | Y | TCGA, GDAC |
Download
|
| Kidney Chromophobe | TCGA | KICH | 113 | Y | TCGA, GDAC |
Download
|
| Kidney renal clear cell carcinoma | TCGA | KIRC | 537 | Y | TCGA, GDAC |
Download
|
| Pan-kidney cohort | TCGA | KIPAN | 941 | Y | TCGA, GDAC |
Download
|
| CPTAC - Lung Adenocarcinoma Confirmatory Study | CPTAC | LUAD | 123 | N | - | - |
| CPTAC - Clear Cell Renal Cell Carcinoma Discovery Cohort | CPTAC | CCRCC | 110 | Y | CPTAC |
Download
|
| CPTAC - Pancreatic Ductal Adenocarcinoma Discovery Study | CPTAC | PDAC | 140 | Y | CPTAC |
Download
|
| Pediatric_Brain_Tumor(CBTTC(The Children Brain Tumor Tissue Consortium)) | CPTAC | BT | 223 | Y | cBioPortal, Publication |
Download
|
| CPTAC - Ovarian Cancer Confirmatory Study | CPTAC | OV | 83 | Y | CPTAC |
Download
|
| CPTAC soft tissue sarcoma study | CPTAC | SARC | 173 | N | - | - |
| CPTAC Gastric Cancer Discover Study | CPTAC | GC | 159 | Y |
Download
|
|
| CPTAC - Uterine Corpus Endometrial Carcinoma Confirmatory Study | CPTAC | UCEC | 138 | Y | Publication |
Download
|
| CPTAC - Lung squamous cell carcinoma | CPTAC | LSCC | 108 | Y |
Download
|
|
| CPTAC - Glioblastoma | CPTAC | GBM | 99 | Y | CPTAC |
Download
|
| CPTAC - Breast cancer prospective cohort | CPTAC | BRCA | 122 | Y | CPTAC |
Download
|
| CPTAC - Uterine Corpus Endometrial Carcinoma | CPTAC | UCEC | 95 | Y | CPTAC |
Download
|
| CPTAC - Lung adenocarcinoma | CPTAC | LUAD | 110 | Y | CPTAC |
Download
|
| CPTAC - Head-and-neck squamous cell carcinoma | CPTAC | HNSCC | 110 | Y | CPTAC |
Download
|
| CPTAC - Colon Adenocarcinoma | CPTAC | COAD | 210 | Y | CPTAC |
Download
|
| CPTAC pancan - Lung adenocarcinoma | CPTAC-pancan | LUAD | 110 | Y | CPTAC, PDC |
Download
|
| CPTAC pancan - Ovarian Cancer | CPTAC-pancan | OV | 83 | Y | CPTAC, PDC |
Download
|
| CPTAC pancan - Uterine Corpus Endometrial Carcinoma | CPTAC-pancan | UCEC | 94 | Y | CPTAC, PDC |
Download
|
| CPTAC pancan - Pancreatic Ductal Adenocarcinoma | CPTAC-pancan | PDAC | 105 | Y | CPTAC, PDC |
Download
|
| CPTAC pancan - Lung squamous cell carcinoma | CPTAC-pancan | LSCC | 108 | Y | CPTAC, PDC |
Download
|
| CPTAC pancan - Head-and-neck squamous cell carcinoma | CPTAC-pancan | HNSCC | 108 | Y | CPTAC, PDC |
Download
|
| CPTAC pancan - Breast cancer prospective cohort | CPTAC-pancan | BRCA | 122 | Y | CPTAC, PDC |
Download
|
| CPTAC pancan - Clear Cell Renal Cell Carcinoma | CPTAC-pancan | CCRCC | 103 | Y | CPTAC, PDC |
Download
|
| CPTAC pancan - Glioblastoma | CPTAC-pancan | GBM | 98 | Y | CPTAC, PDC |
Download
|
| CPTAC pancan - Colon Adenocarcinoma cohort | CPTAC-pancan | COAD | 109 | Y | CPTAC, PDC |
Download
|
| PDX - Breast Cancer | PDX | BRCA | 37 | N | - | - |
| PDX Breast Cancer (BCM_PDX_BRCA 2021) | PDX | BRCA | 94 | N | - | - |
| Breast cancer ER+ PDX | PDX | BRCA | 22 | N | - | - |
| PDX - Breast Cancer (TW) | PDX | BRCA | 16 | N | - | - |
| CellLine Colorectal adenocarcinoma | CL | COADREAD | 44 | N | - | - |
| Brain Metastatic Lung Cancer - Lung adenocarcinoma | BMLC | LUAD | 18 | N | - | - |
| Liverpool - Non-Small Cell Lung Cancer | Liverpool | NSCLC | 194 | N | - | - |
Please Note:
- COADREAD: colorectal, combines COAD + READ
- GBMLGG: glioma, combines GBM + LGG
- KIPAN: pan-kidney, combines KICH + KIRC + KIRP
- STES: stomach-esophogeal, combines STAD + ESCA
(* We acknowledge usage of data from The cancer genome Atlas (TCGA), The Clinical Proteomic Tumor Analysis Consortium (CPTAC) and Broad GDAC)
- Download Manual here Manual version 1.2
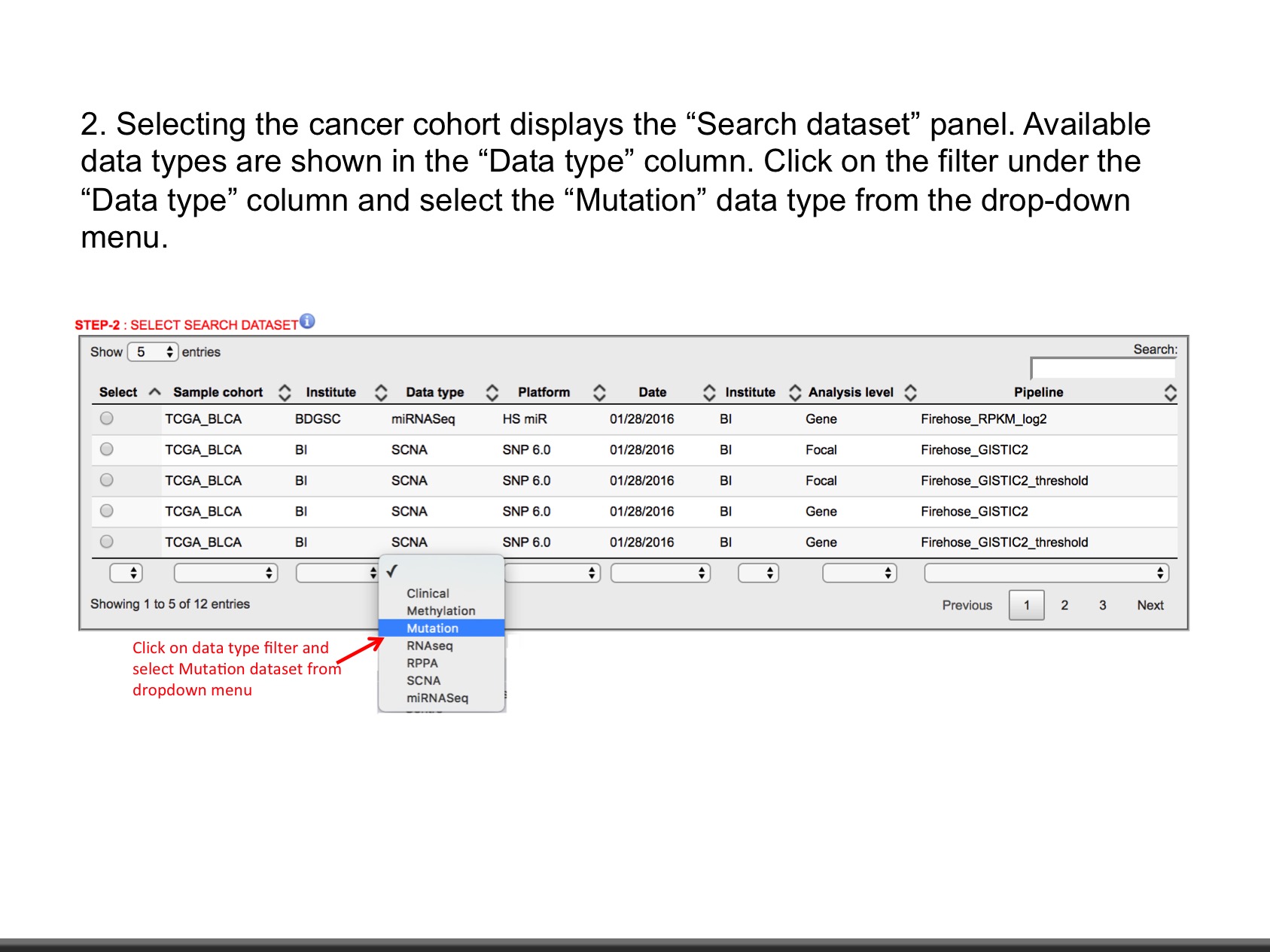
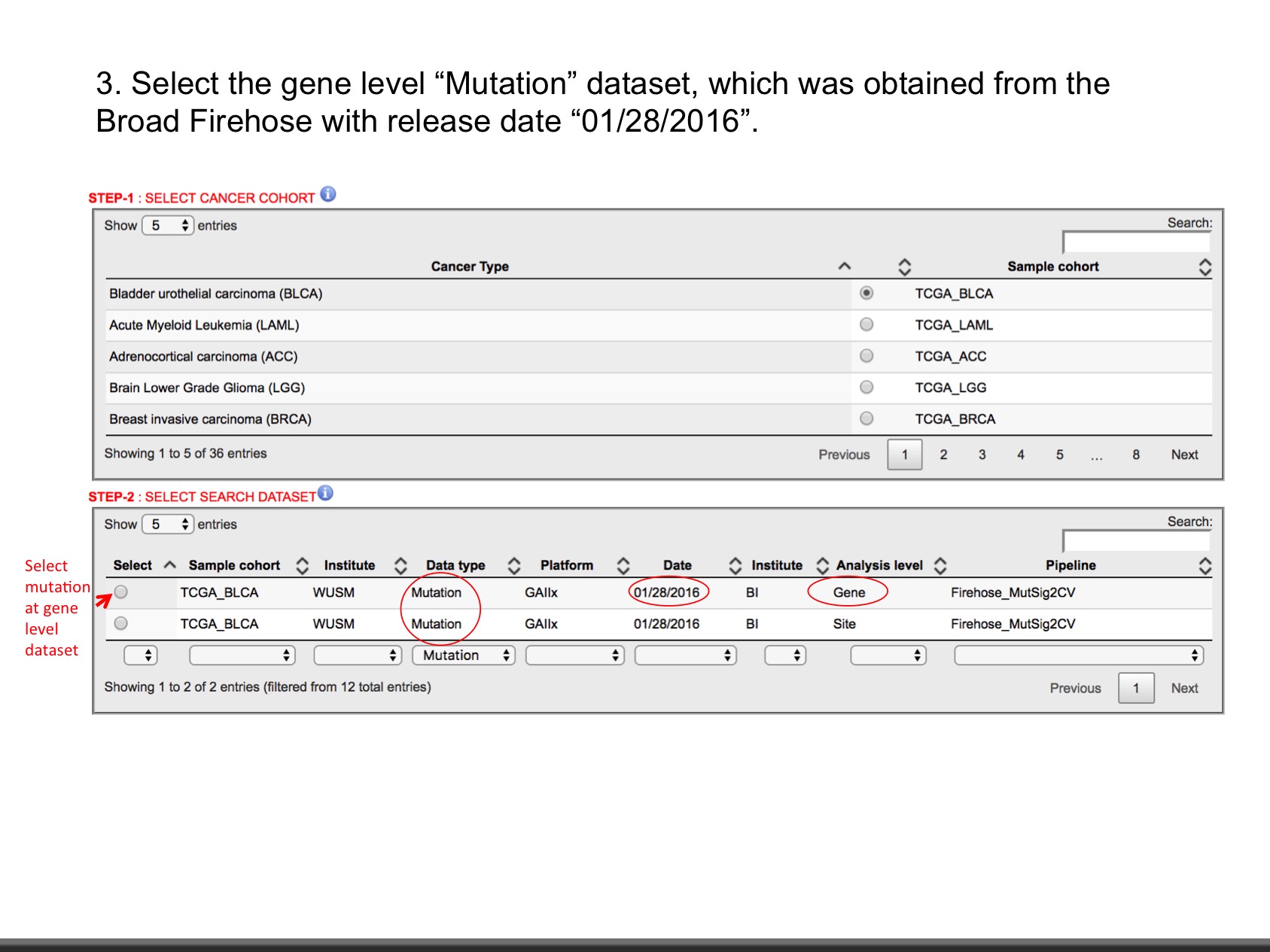
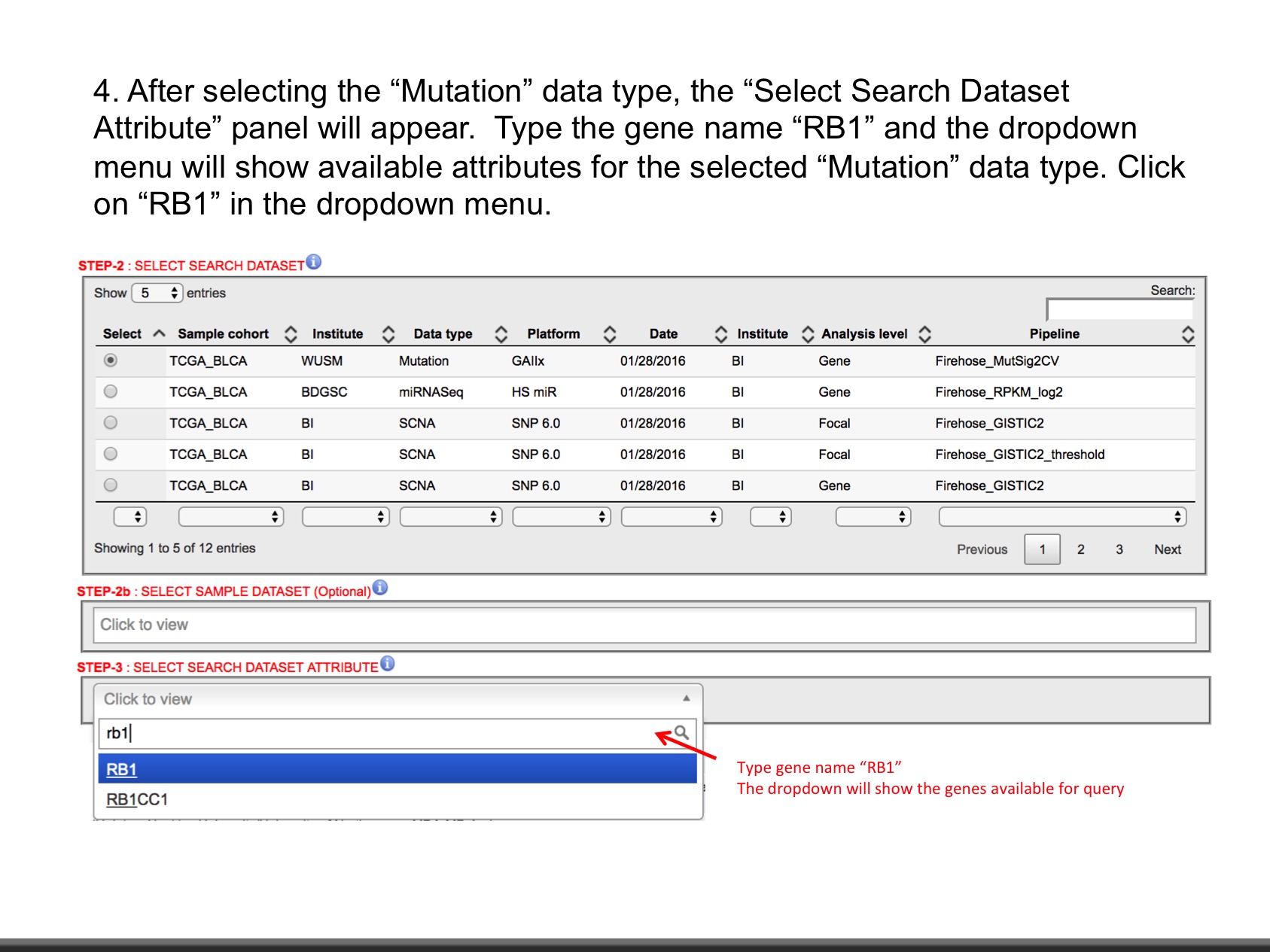
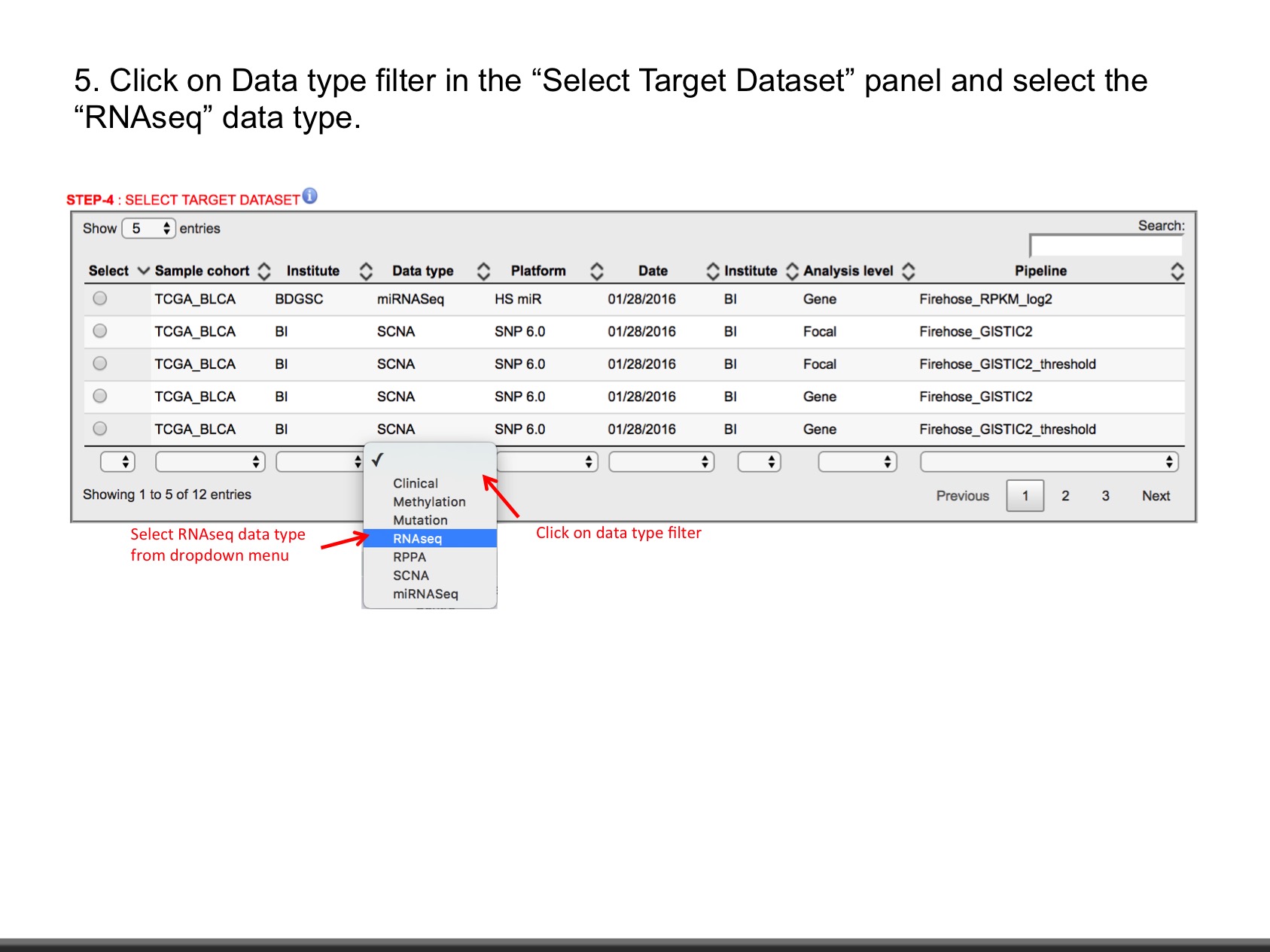
- Explore data as shown in slides.
- Workshop Tutorial Download here
- March 29, 2017 : First version of LinkedOmics portal is available to public.
- Sept 29, 2017 : Workshop at Breast Cancer Center, Baylor College of Medicine. Workshop tutorial can be found here Workshop at BCM
- May 2, 2018 : Workshop at CPTAC3 meeting NIH. Workshop tutorial can be found here Workshop at NIH
- Oct 8, 2018 : WikiPathways summit demo. Workshop tutorial can be found here Summit at UCSF, Gladstone Institute
We are happy to integrate your data into LinkedOmics portal. Follow the instructions given in manual. Please contact admin to share your data (linkedomics.zhanglab@gmail.com).
The tool best run with browser Chrome Version 56.0.2924.87+ (download). If you are using Safari then enable the hidden Develop menu in Safari: Pull down the “Safari” menu and choose “Preferences” Click on the “Advanced” tab. Check the box next to “Show Develop menu in menu bar”. Click Develop menu and select Chrome/Firefox as browser.